Comparing networks#
Comparing networks: Characterizing computational similarity in task-trained recurrent neural networks
By Neuromatch Academy
Content creators: Chris Versteeg
Content reviewers: Chris Versteeg, Hannah Choi, Eva Dyer
Production editors: Konstantine Tsafatinos, Ella Batty, Spiros Chavlis, Samuele Bolotta, Hlib Solodzhuk
Background#
Researchers training artificial networks to perform tasks (e.g., image classification, motor control) have found that the activity in the artificial networks can resemble the activity of biological neurons from brain areas thought to perform similar tasks. Unfortunately, it is unclear whether a superficial similarity in neural activation necessarily translates to a conserved computational strategy. We need ways to assess how well different models are able to capture the computational principles, which will require datasets where the ground-truth computations are known, and we can analyze the similarity between artificial and natural systems. The aim of this project is to explore ways to measure alignment in dynamical systems, and to study different approaches to quantify the changes in representations across different tasks and across different architectures.
Install and import feedback gadget#
Show code cell source
# @title Install and import feedback gadget
!pip install vibecheck datatops --quiet
from vibecheck import DatatopsContentReviewContainer
def content_review(notebook_section: str):
return DatatopsContentReviewContainer(
"", # No text prompt
notebook_section,
{
"url": "https://pmyvdlilci.execute-api.us-east-1.amazonaws.com/klab",
"name": "neuromatch_neuroai",
"user_key": "wb2cxze8",
},
).render()
feedback_prefix = "Project_ComparingNetworks"
Project Background#
Submit your feedback#
Show code cell source
# @title Submit your feedback
content_review(f"{feedback_prefix}_project_background")
Project slides#
If you want to download the slides: https://osf.io/download/vb3tw/
Project Template#
Show code cell source
#@title Project Template
from IPython.display import Image, display
import os
from pathlib import Path
url = "https://github.com/neuromatch/NeuroAI_Course/blob/main/projects/project-notebooks/static/ComputationalSimilarityTemplate.png?raw=true"
display(Image(url=url))

Submit your feedback#
Show code cell source
# @title Submit your feedback
content_review(f"{feedback_prefix}_project_template")
In this notebook, we are going to provide code to get you started on Q1-Q3 of this project!
The basic outline looks like this:
Section 1: Preparing the environment.
Section 2: Overview of the available tasks.
Section 3: Understanding the Three-Bit Flip-Flop task (3BFF).
Section 4: Training a model to perform 3BFF.
Section 5: Inspecting the performance of trained models.
Part 1: Visualizing latent activity
Part 2: Quantifying latent similarity with State R2.
Part 3: Visualizing Fixed-Point architectures.
Section 6: Introduction to Random Target task.
Importantly, we’ve put landmarks in the notebook to indicate:
Interactive exercises
❓❓❓
Cells that will a decent amount of time to run (>5 mins)
⏳⏳⏳
Tutorial links
This project is mostly associated with the materials presented in W1D3, on comparing activities of artificial and biological networks. One of the main techniques here, DSA, completes and empowers your toolbox by enabling dynamic analysis of activity patterns. You will find the Tutorial 2 from the W1D1 the most similar by the model’s architecture and the idea of goal-oriented networks.
Section 1: Preparing the environment#
Disclaimer: As an alternative to Google Colab, Kaggle, and local installation, we have prepared a Dockerfile with the instructions on the virtual environment setup. It will allow you to work locally with no interventions into already installed packages (you will just need to install Docker and run two commands). Grab a Dockerfile, put it in the same folder as this notebook, and follow the instructions in README file.
IF USING COLAB // KAGGLE:
Uncomment the lines of code below and run them in order. The top, the second, and the last cells only need to be run once, but the third cells (envStr) need to be re-run if the Colab // Kaggle notebook crashes. These blocks install the needed dependencies and set up your environment. Notice that the first and third cell contents depend on whether you use Colab or Kaggle.
⏳⏳⏳
Colab // Kaggle installation (Part 1)
Show code cell source
# @markdown Colab // Kaggle installation (Part 1)
# ! git clone https://github.com/neuromatch/ComputationThruDynamicsBenchmark
# %cd ComputationThruDynamicsBenchmark
# ! pip install -e .
## RUN THIS CELL, THEN RESTART SESSION AS PROMPTED (BUTTON AT BOTTOM OF THIS CELL'S FINISHED OUTPUT). DO NOT NEED TO RUN AGAIN
## PLEASE RESTART THE ENVIRONMENT FOR KAGGLE MANUALLY (Run > Restart & clear cell outputs)
Colab // Kaggle installation (Part 2)
Show code cell source
# @markdown Colab // Kaggle installation (Part 2)
# !pip uninstall -y torchaudio torchvision
Colab // Kaggle installation (Part 3)
Show code cell source
# @markdown Colab // Kaggle installation (Part 3)
## GET BACK TO THE DIRECTORY AND CONFIGURE .env
################ COLAB #####################
# %cd /content/ComputationThruDynamicsBenchmark/
# envStr = """HOME_DIR=/content/ComputationThruDynamicsBenchmark/
# #Don't change these
# TRAIN_INPUT_FILE=train_input.h5\nEVAL_INPUT_FILE=eval_input.h5
# EVAL_TARGET_FILE=eval_target.h5
# """
#############################################
################ KAGGLE #####################
# %cd /kaggle/working/ComputationThruDynamicsBenchmark/
# envStr = """HOME_DIR=/kaggle/working/ComputationThruDynamicsBenchmark/
# #Don't change these
# TRAIN_INPUT_FILE=train_input.h5\nEVAL_INPUT_FILE=eval_input.h5
# EVAL_TARGET_FILE=eval_target.h5
# """
##############################################
################ COLAB // KAGGLE #####################
# with open('.env','w') as f:
# f.write(envStr)
##############################################
Colab // Kaggle installation (Part 4)
Show code cell source
# @markdown Colab // Kaggle installation (Part 4)
# !git clone https://github.com/mitchellostrow/DSA
# %cd DSA/
# !pip install -e .
IF RUNNING LOCALLY:
Follow the instructions here to setup the separate environment for this project, or you can run the cell below for general installment.
Local installation
Show code cell source
# @markdown Local installation
import contextlib
import io
import os
dirname = "ComputationThruDynamicsBenchmark"
with contextlib.redirect_stdout(io.StringIO()): #to suppress output
if not os.path.isdir(dirname):
! git clone https://github.com/neuromatch/ComputationThruDynamicsBenchmark
%cd ComputationThruDynamicsBenchmark
! pip install -e .
envStr = """HOME_DIR=ComputationThruDynamicsBenchmark/
#Don't change these
TRAIN_INPUT_FILE=train_input.h5\nEVAL_INPUT_FILE=eval_input.h5
EVAL_TARGET_FILE=eval_target.h5
"""
with open('.env','w') as f:
f.write(envStr)
Submit your feedback#
Show code cell source
# @title Submit your feedback
content_review(f"{feedback_prefix}_preparing_environment")
Section 2: Overview of the available tasks#
First, let’s take a high-level look at the tasks that we are going to use to understand computation in artificial networks!
We’ll start by loading in some packages.
# set the random seed for reproducibility
import random
import dotenv
import pathlib
import os
import logging
# comment the next three lines if you want to see all training logs
pl_loggers = [logging.getLogger(name) for name in logging.root.manager.loggerDict if 'pytorch_lightning' in name]
for pl_log in pl_loggers:
logging.getLogger(pl_log.name).setLevel(logging.WARNING)
random.seed(2024)
dotenv.load_dotenv(override=True)
HOME_DIR = os.getenv("HOME_DIR")
if HOME_DIR is None:
HOME_DIR = ""
print(HOME_DIR)
ComputationThruDynamicsBenchmark/
The Computation-Thru-Dynamics Benchmark has three distinct behavioral tasks.
These tasks are called:
Three-Bit Flip-Flop (3BFF) (see Sussillo & Barak 2013)
MultiTask (See Driscoll et al. 2023)
RandomTarget (See Codol et al. 2023)
We chose these tasks because they represent a variety of task complexities. We have a pretty good understanding of how the simpler tasks operate (3BFF), but really are only starting to scratch the surface of more complex tasks (RandomTarget).
Specificially, in the Random Target task, the actions that the model takes can affect the future inputs, making it an important test case for being able to understand the dynamics of interacting systems!

Each task (which we call a “task environment”) follows a standardized format that allows alternative task environments to be incorporated without any changes to the training pipeline.
Here, we’ll take a walk through the two tasks in the project template (TBFF and RandomTarget) and inspect the behavior of networks trained in these environments.
Submit your feedback#
Show code cell source
# @title Submit your feedback
content_review(f"{feedback_prefix}_overview_of_the_available_tasks")
Section 3: Understanding the Three-Bit Flip-Flop task (3BFF)#
We’re going to start out with the task that launched a thousand Nature papers, the 3-Bit Flip-Flop. Sussillo & Barak 2013 used the three-bit flip-flop in their seminal attempts to understand how dynamics can give rise to computation!
The code snippet below instantiates a “TaskEnv” object, which contains the logic for the NBFF task.
❓❓❓
The default parameters are shown in ./interpretability/task_modeling/configs/env_task/NBFF.yaml, but try changing the parameters below to see how that affects trials generated from the environment. Note that this task is modular in the number of bits as well, so it provides an easy way to scale the dimensionality of a very simple dynamical system.
❓❓❓
from ctd.task_modeling.task_env.task_env import NBitFlipFlop
n = 3 # The number of bits in the flip-flop (default: 3)
trial_length = 500 # The number of time steps in each trial (default: 500)
switch_prob = 0.015 # The probability of an input pulse (default: 0.015 pulses/channel / time step)
noise = 0.15 # The standard deviation of the Gaussian noise added to the input (default: 0.15)
# This line creates the NBitFlipFlop environment. See ctd.task_modeling.task_env.task_env.NBitFlipFlop for more information.
env_3bff = NBitFlipFlop(
n = n,
n_timesteps=trial_length,
switch_prob=switch_prob,
noise=noise
)
# Renders a random trial from the environment
env_3bff.render()
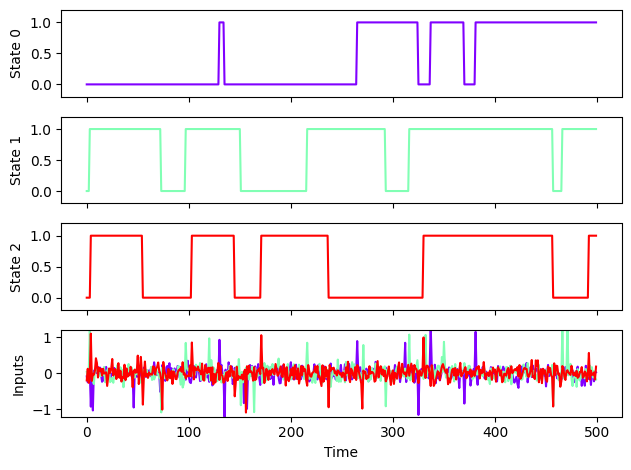
Above, we are plotting the inputs and outputs of the 3BFF task. One trial is 500 time steps, each with a 1% probability of getting an “up” or “down” pulse on each of its 3 input channels. When the task receives an “up” pulse, the state corresponding to that input channel moves from zero to one (if possible), and if a state at one receives a “down” pulse, it goes to zero. In this way, this system acts as 3 bits of memory, encoding 8 potential system states (2^3 states). We add noise to the inputs of the system so that it better reflects realistic computations that a neural circuit might perform.
Try changing the parameters of your 3BFF environment to see how the behavior changes!
Another way to visualize this is to view the three states in 3D. Below, you can see that the 8 potential states appear as the vertices of a cube. Each trial is plotted as a column.
env_3bff.render_3d(n_trials=6)
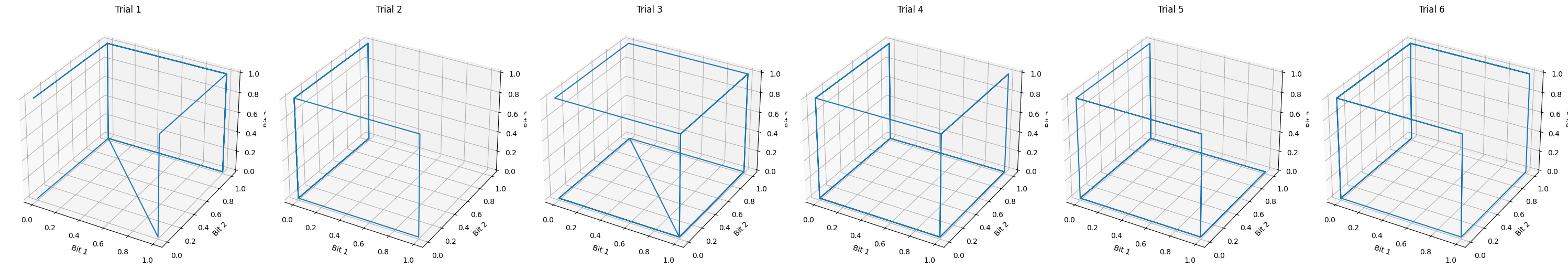
Now that we can see the basic logic of the task, let’s do a basic overview of what task training is!

For task-training, we are simply training a model (e.g., an RNN) to produce a set of outputs given a set of inputs. This input/output relationship defines the task that the model is performing. In the case of 3BFF, an input pulse should cause the model’s output to change in a way that reflects the switching of a bit.
3BFF Training Objective:
3BFF models are trained to minimize the MSE between the desired output and the output of the model, with some other components that pressure the solution to be smooth. If you’re interested in the specifics, the implementation of the loss function can be found as the NBFFLoss object in ctd/task_modeling/task_env/loss_func.py.
Submit your feedback#
Show code cell source
# @title Submit your feedback
content_review(f"{feedback_prefix}_understanding_3bff")
Section 4: Training a model to perform 3BFF#
For this tutorial, we are using PyTorch Lightning to abstract much of the engineering away, allowing you to focus your full attention on the scientific questions you want to tackle!
This segment takes around 8 minutes to train, so I’d recommend planning your runs accordingly!
⏳⏳⏳
The cell below will create a recurrent neural network (RNN) model and use the 3BFF environment to generate samples on which the model will be trained!
Unfortunately, it generates a lot of output, so if you don’t care to see the model progress, set enable_progress_bar to False below.
from ctd.task_modeling.model.rnn import GRU_RNN
from ctd.task_modeling.datamodule.task_datamodule import TaskDataModule
from ctd.task_modeling.task_wrapper.task_wrapper import TaskTrainedWrapper
from pytorch_lightning import Trainer
enable_progress_bar = False
# Step 1: Instantiate the model
rnn = GRU_RNN(latent_size = 128) # Look in ctd/task_modeling/models for alternative choices!
# Step 2: Instantiate the task environment
task_env = env_3bff
# Step 3: Instantiate the task datamodule
task_datamodule = TaskDataModule(task_env, n_samples = 1000, batch_size = 1000)
# Step 4: Instantiate the task wrapper
task_wrapper = TaskTrainedWrapper(learning_rate=1e-3, weight_decay = 1e-8)
# Step 5: Initialize the model with the input and output sizes (3 inputs, 3 outputs, in this case)
rnn.init_model(
input_size = task_env.observation_space.shape[0],
output_size = task_env.action_space.shape[0]
)
# Step 6: Set the environment and model in the task wrapper
task_wrapper.set_environment(task_env)
task_wrapper.set_model(rnn)
# Step 7: Define the PyTorch Lightning Trainer object
trainer = Trainer(max_epochs=500, enable_progress_bar=enable_progress_bar)
# Step 8: Fit the model
trainer.fit(task_wrapper, task_datamodule)
---------------------------------------------------------------------------
TypeError Traceback (most recent call last)
Cell In[18], line 34
31 trainer = Trainer(max_epochs=500, enable_progress_bar=enable_progress_bar)
33 # Step 8: Fit the model
---> 34 trainer.fit(task_wrapper, task_datamodule)
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/pytorch_lightning/trainer/trainer.py:544, in Trainer.fit(self, model, train_dataloaders, val_dataloaders, datamodule, ckpt_path)
542 self.state.status = TrainerStatus.RUNNING
543 self.training = True
--> 544 call._call_and_handle_interrupt(
545 self, self._fit_impl, model, train_dataloaders, val_dataloaders, datamodule, ckpt_path
546 )
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/pytorch_lightning/trainer/call.py:44, in _call_and_handle_interrupt(trainer, trainer_fn, *args, **kwargs)
42 if trainer.strategy.launcher is not None:
43 return trainer.strategy.launcher.launch(trainer_fn, *args, trainer=trainer, **kwargs)
---> 44 return trainer_fn(*args, **kwargs)
46 except _TunerExitException:
47 _call_teardown_hook(trainer)
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/pytorch_lightning/trainer/trainer.py:580, in Trainer._fit_impl(self, model, train_dataloaders, val_dataloaders, datamodule, ckpt_path)
573 assert self.state.fn is not None
574 ckpt_path = self._checkpoint_connector._select_ckpt_path(
575 self.state.fn,
576 ckpt_path,
577 model_provided=True,
578 model_connected=self.lightning_module is not None,
579 )
--> 580 self._run(model, ckpt_path=ckpt_path)
582 assert self.state.stopped
583 self.training = False
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/pytorch_lightning/trainer/trainer.py:950, in Trainer._run(self, model, ckpt_path)
947 self.strategy.setup_environment()
948 self.__setup_profiler()
--> 950 call._call_setup_hook(self) # allow user to setup lightning_module in accelerator environment
952 # check if we should delay restoring checkpoint till later
953 if not self.strategy.restore_checkpoint_after_setup:
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/pytorch_lightning/trainer/call.py:86, in _call_setup_hook(trainer)
84 # Trigger lazy creation of experiment in loggers so loggers have their metadata available
85 for logger in trainer.loggers:
---> 86 if hasattr(logger, "experiment"):
87 _ = logger.experiment
89 trainer.strategy.barrier("pre_setup")
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/lightning_fabric/loggers/logger.py:118, in rank_zero_experiment.<locals>.experiment(self)
116 if rank_zero_only.rank > 0:
117 return _DummyExperiment()
--> 118 return fn(self)
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/lightning_fabric/loggers/tensorboard.py:187, in TensorBoardLogger.experiment(self)
184 self._fs.makedirs(self.root_dir, exist_ok=True)
186 if _TENSORBOARD_AVAILABLE:
--> 187 from torch.utils.tensorboard import SummaryWriter
188 else:
189 from tensorboardX import SummaryWriter # type: ignore[no-redef]
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/torch/utils/tensorboard/__init__.py:12
9 del Version
10 del tensorboard
---> 12 from .writer import FileWriter, SummaryWriter # noqa: F401
13 from tensorboard.summary.writer.record_writer import RecordWriter # noqa: F401
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/torch/utils/tensorboard/writer.py:10
7 import torch
9 from tensorboard.compat import tf
---> 10 from tensorboard.compat.proto import event_pb2
11 from tensorboard.compat.proto.event_pb2 import Event, SessionLog
12 from tensorboard.plugins.projector.projector_config_pb2 import ProjectorConfig
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/tensorboard/compat/proto/event_pb2.py:17
12 # @@protoc_insertion_point(imports)
14 _sym_db = _symbol_database.Default()
---> 17 from tensorboard.compat.proto import summary_pb2 as tensorboard_dot_compat_dot_proto_dot_summary__pb2
20 DESCRIPTOR = _descriptor.FileDescriptor(
21 name='tensorboard/compat/proto/event.proto',
22 package='tensorboard',
(...)
26 ,
27 dependencies=[tensorboard_dot_compat_dot_proto_dot_summary__pb2.DESCRIPTOR,])
29 _WORKERHEALTH = _descriptor.EnumDescriptor(
30 name='WorkerHealth',
31 full_name='tensorboard.WorkerHealth',
(...)
55 serialized_end=1319,
56 )
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/tensorboard/compat/proto/summary_pb2.py:17
12 # @@protoc_insertion_point(imports)
14 _sym_db = _symbol_database.Default()
---> 17 from tensorboard.compat.proto import tensor_pb2 as tensorboard_dot_compat_dot_proto_dot_tensor__pb2
20 DESCRIPTOR = _descriptor.FileDescriptor(
21 name='tensorboard/compat/proto/summary.proto',
22 package='tensorboard',
(...)
26 ,
27 dependencies=[tensorboard_dot_compat_dot_proto_dot_tensor__pb2.DESCRIPTOR,])
29 _DATACLASS = _descriptor.EnumDescriptor(
30 name='DataClass',
31 full_name='tensorboard.DataClass',
(...)
55 serialized_end=1228,
56 )
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/tensorboard/compat/proto/tensor_pb2.py:16
11 # @@protoc_insertion_point(imports)
13 _sym_db = _symbol_database.Default()
---> 16 from tensorboard.compat.proto import resource_handle_pb2 as tensorboard_dot_compat_dot_proto_dot_resource__handle__pb2
17 from tensorboard.compat.proto import tensor_shape_pb2 as tensorboard_dot_compat_dot_proto_dot_tensor__shape__pb2
18 from tensorboard.compat.proto import types_pb2 as tensorboard_dot_compat_dot_proto_dot_types__pb2
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/tensorboard/compat/proto/resource_handle_pb2.py:16
11 # @@protoc_insertion_point(imports)
13 _sym_db = _symbol_database.Default()
---> 16 from tensorboard.compat.proto import tensor_shape_pb2 as tensorboard_dot_compat_dot_proto_dot_tensor__shape__pb2
17 from tensorboard.compat.proto import types_pb2 as tensorboard_dot_compat_dot_proto_dot_types__pb2
20 DESCRIPTOR = _descriptor.FileDescriptor(
21 name='tensorboard/compat/proto/resource_handle.proto',
22 package='tensorboard',
(...)
26 ,
27 dependencies=[tensorboard_dot_compat_dot_proto_dot_tensor__shape__pb2.DESCRIPTOR,tensorboard_dot_compat_dot_proto_dot_types__pb2.DESCRIPTOR,])
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/tensorboard/compat/proto/tensor_shape_pb2.py:36
13 _sym_db = _symbol_database.Default()
18 DESCRIPTOR = _descriptor.FileDescriptor(
19 name='tensorboard/compat/proto/tensor_shape.proto',
20 package='tensorboard',
(...)
23 serialized_pb=_b('\n+tensorboard/compat/proto/tensor_shape.proto\x12\x0btensorboard\"{\n\x10TensorShapeProto\x12.\n\x03\x64im\x18\x02 \x03(\x0b\x32!.tensorboard.TensorShapeProto.Dim\x12\x14\n\x0cunknown_rank\x18\x03 \x01(\x08\x1a!\n\x03\x44im\x12\x0c\n\x04size\x18\x01 \x01(\x03\x12\x0c\n\x04name\x18\x02 \x01(\tB\x87\x01\n\x18org.tensorflow.frameworkB\x11TensorShapeProtosP\x01ZSgithub.com/tensorflow/tensorflow/tensorflow/go/core/framework/tensor_shape_go_proto\xf8\x01\x01\x62\x06proto3')
24 )
29 _TENSORSHAPEPROTO_DIM = _descriptor.Descriptor(
30 name='Dim',
31 full_name='tensorboard.TensorShapeProto.Dim',
32 filename=None,
33 file=DESCRIPTOR,
34 containing_type=None,
35 fields=[
---> 36 _descriptor.FieldDescriptor(
37 name='size', full_name='tensorboard.TensorShapeProto.Dim.size', index=0,
38 number=1, type=3, cpp_type=2, label=1,
39 has_default_value=False, default_value=0,
40 message_type=None, enum_type=None, containing_type=None,
41 is_extension=False, extension_scope=None,
42 serialized_options=None, file=DESCRIPTOR),
43 _descriptor.FieldDescriptor(
44 name='name', full_name='tensorboard.TensorShapeProto.Dim.name', index=1,
45 number=2, type=9, cpp_type=9, label=1,
46 has_default_value=False, default_value=_b("").decode('utf-8'),
47 message_type=None, enum_type=None, containing_type=None,
48 is_extension=False, extension_scope=None,
49 serialized_options=None, file=DESCRIPTOR),
50 ],
51 extensions=[
52 ],
53 nested_types=[],
54 enum_types=[
55 ],
56 serialized_options=None,
57 is_extendable=False,
58 syntax='proto3',
59 extension_ranges=[],
60 oneofs=[
61 ],
62 serialized_start=150,
63 serialized_end=183,
64 )
66 _TENSORSHAPEPROTO = _descriptor.Descriptor(
67 name='TensorShapeProto',
68 full_name='tensorboard.TensorShapeProto',
(...)
100 serialized_end=183,
101 )
103 _TENSORSHAPEPROTO_DIM.containing_type = _TENSORSHAPEPROTO
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/google/protobuf/descriptor.py:675, in FieldDescriptor.__new__(cls, name, full_name, index, number, type, cpp_type, label, default_value, message_type, enum_type, containing_type, is_extension, extension_scope, options, serialized_options, has_default_value, containing_oneof, json_name, file, create_key)
652 def __new__(
653 cls,
654 name,
(...)
673 create_key=None,
674 ): # pylint: disable=redefined-builtin
--> 675 _message.Message._CheckCalledFromGeneratedFile()
676 if is_extension:
677 return _message.default_pool.FindExtensionByName(full_name)
TypeError: Descriptors cannot be created directly.
If this call came from a _pb2.py file, your generated code is out of date and must be regenerated with protoc >= 3.19.0.
If you cannot immediately regenerate your protos, some other possible workarounds are:
1. Downgrade the protobuf package to 3.20.x or lower.
2. Set PROTOCOL_BUFFERS_PYTHON_IMPLEMENTATION=python (but this will use pure-Python parsing and will be much slower).
More information: https://developers.google.com/protocol-buffers/docs/news/2022-05-06#python-updates
Now, we use pickle to save the trained model and datamodule for future analyses!
❓❓❓
Once you get this model trained, feel free to try changing the hyperparameters to see if you can get the model to train faster!
❓❓❓
import pickle
# save model as .pkl
save_dir = pathlib.Path(HOME_DIR) / "models_GRU_128"
save_dir.mkdir(exist_ok=True)
with open(save_dir / "model.pkl", "wb") as f:
pickle.dump(task_wrapper, f)
# save datamodule as .pkl
with open(save_dir / "datamodule_sim.pkl", "wb") as f:
pickle.dump(task_datamodule, f)
So that we can start comparing our models, we’re going to train a second GRU_RNN to perform the 3BFF task, except this time, we’ll use an alternative model called a Neural ODE!
Notice that we’re using the same datamodule as for the first model, meaning that we can directly compare the two models trial-by-trial.
Again, this will take around 10 minutes to train!
⏳⏳⏳
from ctd.task_modeling.model.node import NODE
enable_progress_bar = False
rnn = NODE(latent_size = 3, num_layers = 3, layer_hidden_size=64) # Look in ctd/task_modeling/models for alternative choices!
task_wrapper = TaskTrainedWrapper(learning_rate=1e-3, weight_decay = 1e-10)
rnn.init_model(
input_size = task_env.observation_space.shape[0],
output_size = task_env.action_space.shape[0]
)
task_wrapper.set_environment(task_env)
task_wrapper.set_model(rnn)
trainer = Trainer(max_epochs=500, enable_progress_bar=enable_progress_bar)
trainer.fit(task_wrapper, task_datamodule)
save_dir = pathlib.Path(HOME_DIR) / "models_NODE_3"
save_dir.mkdir(exist_ok=True)
with open(save_dir / "model.pkl", "wb") as f:
pickle.dump(task_wrapper, f)
# save datamodule as .pkl
with open(save_dir / "datamodule_sim.pkl", "wb") as f:
pickle.dump(task_datamodule, f)
---------------------------------------------------------------------------
TypeError Traceback (most recent call last)
Cell In[20], line 13
10 task_wrapper.set_model(rnn)
11 trainer = Trainer(max_epochs=500, enable_progress_bar=enable_progress_bar)
---> 13 trainer.fit(task_wrapper, task_datamodule)
15 save_dir = pathlib.Path(HOME_DIR) / "models_NODE_3"
16 save_dir.mkdir(exist_ok=True)
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/pytorch_lightning/trainer/trainer.py:544, in Trainer.fit(self, model, train_dataloaders, val_dataloaders, datamodule, ckpt_path)
542 self.state.status = TrainerStatus.RUNNING
543 self.training = True
--> 544 call._call_and_handle_interrupt(
545 self, self._fit_impl, model, train_dataloaders, val_dataloaders, datamodule, ckpt_path
546 )
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/pytorch_lightning/trainer/call.py:44, in _call_and_handle_interrupt(trainer, trainer_fn, *args, **kwargs)
42 if trainer.strategy.launcher is not None:
43 return trainer.strategy.launcher.launch(trainer_fn, *args, trainer=trainer, **kwargs)
---> 44 return trainer_fn(*args, **kwargs)
46 except _TunerExitException:
47 _call_teardown_hook(trainer)
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/pytorch_lightning/trainer/trainer.py:580, in Trainer._fit_impl(self, model, train_dataloaders, val_dataloaders, datamodule, ckpt_path)
573 assert self.state.fn is not None
574 ckpt_path = self._checkpoint_connector._select_ckpt_path(
575 self.state.fn,
576 ckpt_path,
577 model_provided=True,
578 model_connected=self.lightning_module is not None,
579 )
--> 580 self._run(model, ckpt_path=ckpt_path)
582 assert self.state.stopped
583 self.training = False
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/pytorch_lightning/trainer/trainer.py:950, in Trainer._run(self, model, ckpt_path)
947 self.strategy.setup_environment()
948 self.__setup_profiler()
--> 950 call._call_setup_hook(self) # allow user to setup lightning_module in accelerator environment
952 # check if we should delay restoring checkpoint till later
953 if not self.strategy.restore_checkpoint_after_setup:
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/pytorch_lightning/trainer/call.py:86, in _call_setup_hook(trainer)
84 # Trigger lazy creation of experiment in loggers so loggers have their metadata available
85 for logger in trainer.loggers:
---> 86 if hasattr(logger, "experiment"):
87 _ = logger.experiment
89 trainer.strategy.barrier("pre_setup")
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/lightning_fabric/loggers/logger.py:118, in rank_zero_experiment.<locals>.experiment(self)
116 if rank_zero_only.rank > 0:
117 return _DummyExperiment()
--> 118 return fn(self)
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/lightning_fabric/loggers/tensorboard.py:187, in TensorBoardLogger.experiment(self)
184 self._fs.makedirs(self.root_dir, exist_ok=True)
186 if _TENSORBOARD_AVAILABLE:
--> 187 from torch.utils.tensorboard import SummaryWriter
188 else:
189 from tensorboardX import SummaryWriter # type: ignore[no-redef]
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/torch/utils/tensorboard/__init__.py:12
9 del Version
10 del tensorboard
---> 12 from .writer import FileWriter, SummaryWriter # noqa: F401
13 from tensorboard.summary.writer.record_writer import RecordWriter # noqa: F401
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/torch/utils/tensorboard/writer.py:10
7 import torch
9 from tensorboard.compat import tf
---> 10 from tensorboard.compat.proto import event_pb2
11 from tensorboard.compat.proto.event_pb2 import Event, SessionLog
12 from tensorboard.plugins.projector.projector_config_pb2 import ProjectorConfig
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/tensorboard/compat/proto/event_pb2.py:17
12 # @@protoc_insertion_point(imports)
14 _sym_db = _symbol_database.Default()
---> 17 from tensorboard.compat.proto import summary_pb2 as tensorboard_dot_compat_dot_proto_dot_summary__pb2
20 DESCRIPTOR = _descriptor.FileDescriptor(
21 name='tensorboard/compat/proto/event.proto',
22 package='tensorboard',
(...)
26 ,
27 dependencies=[tensorboard_dot_compat_dot_proto_dot_summary__pb2.DESCRIPTOR,])
29 _WORKERHEALTH = _descriptor.EnumDescriptor(
30 name='WorkerHealth',
31 full_name='tensorboard.WorkerHealth',
(...)
55 serialized_end=1319,
56 )
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/tensorboard/compat/proto/summary_pb2.py:17
12 # @@protoc_insertion_point(imports)
14 _sym_db = _symbol_database.Default()
---> 17 from tensorboard.compat.proto import tensor_pb2 as tensorboard_dot_compat_dot_proto_dot_tensor__pb2
20 DESCRIPTOR = _descriptor.FileDescriptor(
21 name='tensorboard/compat/proto/summary.proto',
22 package='tensorboard',
(...)
26 ,
27 dependencies=[tensorboard_dot_compat_dot_proto_dot_tensor__pb2.DESCRIPTOR,])
29 _DATACLASS = _descriptor.EnumDescriptor(
30 name='DataClass',
31 full_name='tensorboard.DataClass',
(...)
55 serialized_end=1228,
56 )
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/tensorboard/compat/proto/tensor_pb2.py:16
11 # @@protoc_insertion_point(imports)
13 _sym_db = _symbol_database.Default()
---> 16 from tensorboard.compat.proto import resource_handle_pb2 as tensorboard_dot_compat_dot_proto_dot_resource__handle__pb2
17 from tensorboard.compat.proto import tensor_shape_pb2 as tensorboard_dot_compat_dot_proto_dot_tensor__shape__pb2
18 from tensorboard.compat.proto import types_pb2 as tensorboard_dot_compat_dot_proto_dot_types__pb2
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/tensorboard/compat/proto/resource_handle_pb2.py:16
11 # @@protoc_insertion_point(imports)
13 _sym_db = _symbol_database.Default()
---> 16 from tensorboard.compat.proto import tensor_shape_pb2 as tensorboard_dot_compat_dot_proto_dot_tensor__shape__pb2
17 from tensorboard.compat.proto import types_pb2 as tensorboard_dot_compat_dot_proto_dot_types__pb2
20 DESCRIPTOR = _descriptor.FileDescriptor(
21 name='tensorboard/compat/proto/resource_handle.proto',
22 package='tensorboard',
(...)
26 ,
27 dependencies=[tensorboard_dot_compat_dot_proto_dot_tensor__shape__pb2.DESCRIPTOR,tensorboard_dot_compat_dot_proto_dot_types__pb2.DESCRIPTOR,])
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/tensorboard/compat/proto/tensor_shape_pb2.py:36
13 _sym_db = _symbol_database.Default()
18 DESCRIPTOR = _descriptor.FileDescriptor(
19 name='tensorboard/compat/proto/tensor_shape.proto',
20 package='tensorboard',
(...)
23 serialized_pb=_b('\n+tensorboard/compat/proto/tensor_shape.proto\x12\x0btensorboard\"{\n\x10TensorShapeProto\x12.\n\x03\x64im\x18\x02 \x03(\x0b\x32!.tensorboard.TensorShapeProto.Dim\x12\x14\n\x0cunknown_rank\x18\x03 \x01(\x08\x1a!\n\x03\x44im\x12\x0c\n\x04size\x18\x01 \x01(\x03\x12\x0c\n\x04name\x18\x02 \x01(\tB\x87\x01\n\x18org.tensorflow.frameworkB\x11TensorShapeProtosP\x01ZSgithub.com/tensorflow/tensorflow/tensorflow/go/core/framework/tensor_shape_go_proto\xf8\x01\x01\x62\x06proto3')
24 )
29 _TENSORSHAPEPROTO_DIM = _descriptor.Descriptor(
30 name='Dim',
31 full_name='tensorboard.TensorShapeProto.Dim',
32 filename=None,
33 file=DESCRIPTOR,
34 containing_type=None,
35 fields=[
---> 36 _descriptor.FieldDescriptor(
37 name='size', full_name='tensorboard.TensorShapeProto.Dim.size', index=0,
38 number=1, type=3, cpp_type=2, label=1,
39 has_default_value=False, default_value=0,
40 message_type=None, enum_type=None, containing_type=None,
41 is_extension=False, extension_scope=None,
42 serialized_options=None, file=DESCRIPTOR),
43 _descriptor.FieldDescriptor(
44 name='name', full_name='tensorboard.TensorShapeProto.Dim.name', index=1,
45 number=2, type=9, cpp_type=9, label=1,
46 has_default_value=False, default_value=_b("").decode('utf-8'),
47 message_type=None, enum_type=None, containing_type=None,
48 is_extension=False, extension_scope=None,
49 serialized_options=None, file=DESCRIPTOR),
50 ],
51 extensions=[
52 ],
53 nested_types=[],
54 enum_types=[
55 ],
56 serialized_options=None,
57 is_extendable=False,
58 syntax='proto3',
59 extension_ranges=[],
60 oneofs=[
61 ],
62 serialized_start=150,
63 serialized_end=183,
64 )
66 _TENSORSHAPEPROTO = _descriptor.Descriptor(
67 name='TensorShapeProto',
68 full_name='tensorboard.TensorShapeProto',
(...)
100 serialized_end=183,
101 )
103 _TENSORSHAPEPROTO_DIM.containing_type = _TENSORSHAPEPROTO
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/google/protobuf/descriptor.py:675, in FieldDescriptor.__new__(cls, name, full_name, index, number, type, cpp_type, label, default_value, message_type, enum_type, containing_type, is_extension, extension_scope, options, serialized_options, has_default_value, containing_oneof, json_name, file, create_key)
652 def __new__(
653 cls,
654 name,
(...)
673 create_key=None,
674 ): # pylint: disable=redefined-builtin
--> 675 _message.Message._CheckCalledFromGeneratedFile()
676 if is_extension:
677 return _message.default_pool.FindExtensionByName(full_name)
TypeError: Descriptors cannot be created directly.
If this call came from a _pb2.py file, your generated code is out of date and must be regenerated with protoc >= 3.19.0.
If you cannot immediately regenerate your protos, some other possible workarounds are:
1. Downgrade the protobuf package to 3.20.x or lower.
2. Set PROTOCOL_BUFFERS_PYTHON_IMPLEMENTATION=python (but this will use pure-Python parsing and will be much slower).
More information: https://developers.google.com/protocol-buffers/docs/news/2022-05-06#python-updates
Submit your feedback#
Show code cell source
# @title Submit your feedback
content_review(f"{feedback_prefix}_training_a_model_to_perform_3bff")
Section 5: Inspecting the performance of trained models#
Now that the models have been trained, let’s see if we can determine how similar their computational strategies are to each other!
To make your life easier, we’ve provided an “analysis” object that abstracts away much of the data handling, allowing you to work more easily with the data from the models.
The analysis object also offers visualization tools that can help to see how well the trained model learned to perform the task!
For example, plot_trial_io is a function that plots (for a specified number of trials):
Latent activity
Controlled output
Target output
Noisy inputs to model
❓❓❓
Try changing trials that are plotted. Do the models capture all of the states equally well?
❓❓❓
Part 1: Visualizing latent activity#
from ctd.comparison.analysis.tt.tt import Analysis_TT
fpath_GRU_128 = HOME_DIR + "models_GRU_128/"
# Create the analysis object:
analysis_GRU_128 = Analysis_TT(
run_name = "GRU_128_3bff",
filepath = fpath_GRU_128)
analysis_GRU_128.plot_trial_io(num_trials = 2)
---------------------------------------------------------------------------
TypeError Traceback (most recent call last)
Cell In[22], line 1
----> 1 from ctd.comparison.analysis.tt.tt import Analysis_TT
3 fpath_GRU_128 = HOME_DIR + "models_GRU_128/"
4 # Create the analysis object:
File ~/work/NeuroAI_Course/NeuroAI_Course/projects/project-notebooks/ComputationThruDynamicsBenchmark/ctd/comparison/analysis/tt/tt.py:9
7 import numpy as np
8 import torch
----> 9 from DSA.stats import dsa_bw_data_splits, dsa_to_id
10 from sklearn.decomposition import PCA
12 from ctd.comparison.analysis.analysis import Analysis
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/DSA/__init__.py:3
1 __version__ = "2.0.0"
----> 3 from DSA.dsa import DSA, ControllabilitySimilarityTransformDistConfig, GeneralizedDSA, InputDSA, SimilarityTransformDistConfig
4 from DSA.dsa import DefaultDMDConfig as DMDConfig
5 from DSA.dsa import pyKoopmanDMDConfig, SubspaceDMDcConfig, DMDcConfig
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/DSA/dsa.py:5
3 from DSA.dmdc import DMDc as DefaultDMDc
4 from DSA.subspace_dmdc import SubspaceDMDc
----> 5 from DSA.simdist import SimilarityTransformDist
6 from typing import Literal
7 import torch
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/DSA/simdist.py:9
6 import torch.nn.utils.parametrize as parametrize
7 import warnings
----> 9 from ot import dist, emd, emd2, sinkhorn2
11 try:
12 from .dmd import DMD
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/ot/__init__.py:20
1 """
2 .. warning::
3 The list of automatically imported sub-modules is as follows:
(...)
11 - :any:`ot.plot` : depends on :code:`matplotlib`
12 """
14 # Author: Remi Flamary <remi.flamary@unice.fr>
15 # Nicolas Courty <ncourty@irisa.fr>
16 #
17 # License: MIT License
18
19 # All submodules and packages
---> 20 from . import lp
21 from . import bregman
22 from . import optim
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/ot/lp/__init__.py:11
2 """
3 Solvers for the original linear program OT problem.
4
5 """
7 # Author: Remi Flamary <remi.flamary@unice.fr>
8 #
9 # License: MIT License
---> 11 from .dmmot import dmmot_monge_1dgrid_loss, dmmot_monge_1dgrid_optimize
12 from ._network_simplex import emd, emd2
13 from ._barycenter_solvers import (
14 barycenter,
15 free_support_barycenter,
(...)
19 NorthWestMMGluing,
20 )
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/ot/lp/dmmot.py:12
6 # Author: Ronak Mehta <ronakrm@cs.wisc.edu>
7 # Xizheng Yu <xyu354@wisc.edu>
8 #
9 # License: MIT License
11 import numpy as np
---> 12 from ..backend import get_backend
15 def dist_monge_max_min(i):
16 r"""
17 A tensor :math:c is Monge if for all valid :math:i_1, \ldots i_d and
18 :math:j_1, \ldots, j_d,
(...)
54 Workshop on Discrete Algorithms.
55 """
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/ot/backend.py:148
146 if not os.environ.get(DISABLE_TF_KEY, False):
147 try:
--> 148 import tensorflow as tf
149 import tensorflow.experimental.numpy as tnp
151 tf_type = tf.Tensor
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/tensorflow/__init__.py:37
34 import sys as _sys
35 import typing as _typing
---> 37 from tensorflow.python.tools import module_util as _module_util
38 from tensorflow.python.util.lazy_loader import LazyLoader as _LazyLoader
40 # Make sure code inside the TensorFlow codebase can use tf2.enabled() at import.
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/tensorflow/python/__init__.py:37
29 # We aim to keep this file minimal and ideally remove completely.
30 # If you are adding a new file with @tf_export decorators,
31 # import it in modules_with_exports.py instead.
32
33 # go/tf-wildcard-import
34 # pylint: disable=wildcard-import,g-bad-import-order,g-import-not-at-top
36 from tensorflow.python import pywrap_tensorflow as _pywrap_tensorflow
---> 37 from tensorflow.python.eager import context
39 # pylint: enable=wildcard-import
40
41 # Bring in subpackages.
42 from tensorflow.python import data
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/tensorflow/python/eager/context.py:29
26 import numpy as np
27 import six
---> 29 from tensorflow.core.framework import function_pb2
30 from tensorflow.core.protobuf import config_pb2
31 from tensorflow.core.protobuf import coordination_config_pb2
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/tensorflow/core/framework/function_pb2.py:16
11 # @@protoc_insertion_point(imports)
13 _sym_db = _symbol_database.Default()
---> 16 from tensorflow.core.framework import attr_value_pb2 as tensorflow_dot_core_dot_framework_dot_attr__value__pb2
17 from tensorflow.core.framework import node_def_pb2 as tensorflow_dot_core_dot_framework_dot_node__def__pb2
18 from tensorflow.core.framework import op_def_pb2 as tensorflow_dot_core_dot_framework_dot_op__def__pb2
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/tensorflow/core/framework/attr_value_pb2.py:16
11 # @@protoc_insertion_point(imports)
13 _sym_db = _symbol_database.Default()
---> 16 from tensorflow.core.framework import tensor_pb2 as tensorflow_dot_core_dot_framework_dot_tensor__pb2
17 from tensorflow.core.framework import tensor_shape_pb2 as tensorflow_dot_core_dot_framework_dot_tensor__shape__pb2
18 from tensorflow.core.framework import types_pb2 as tensorflow_dot_core_dot_framework_dot_types__pb2
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/tensorflow/core/framework/tensor_pb2.py:16
11 # @@protoc_insertion_point(imports)
13 _sym_db = _symbol_database.Default()
---> 16 from tensorflow.core.framework import resource_handle_pb2 as tensorflow_dot_core_dot_framework_dot_resource__handle__pb2
17 from tensorflow.core.framework import tensor_shape_pb2 as tensorflow_dot_core_dot_framework_dot_tensor__shape__pb2
18 from tensorflow.core.framework import types_pb2 as tensorflow_dot_core_dot_framework_dot_types__pb2
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/tensorflow/core/framework/resource_handle_pb2.py:16
11 # @@protoc_insertion_point(imports)
13 _sym_db = _symbol_database.Default()
---> 16 from tensorflow.core.framework import tensor_shape_pb2 as tensorflow_dot_core_dot_framework_dot_tensor__shape__pb2
17 from tensorflow.core.framework import types_pb2 as tensorflow_dot_core_dot_framework_dot_types__pb2
20 DESCRIPTOR = _descriptor.FileDescriptor(
21 name='tensorflow/core/framework/resource_handle.proto',
22 package='tensorflow',
(...)
26 ,
27 dependencies=[tensorflow_dot_core_dot_framework_dot_tensor__shape__pb2.DESCRIPTOR,tensorflow_dot_core_dot_framework_dot_types__pb2.DESCRIPTOR,])
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/tensorflow/core/framework/tensor_shape_pb2.py:36
13 _sym_db = _symbol_database.Default()
18 DESCRIPTOR = _descriptor.FileDescriptor(
19 name='tensorflow/core/framework/tensor_shape.proto',
20 package='tensorflow',
(...)
23 serialized_pb=_b('\n,tensorflow/core/framework/tensor_shape.proto\x12\ntensorflow\"z\n\x10TensorShapeProto\x12-\n\x03\x64im\x18\x02 \x03(\x0b\x32 .tensorflow.TensorShapeProto.Dim\x12\x14\n\x0cunknown_rank\x18\x03 \x01(\x08\x1a!\n\x03\x44im\x12\x0c\n\x04size\x18\x01 \x01(\x03\x12\x0c\n\x04name\x18\x02 \x01(\tB\x87\x01\n\x18org.tensorflow.frameworkB\x11TensorShapeProtosP\x01ZSgithub.com/tensorflow/tensorflow/tensorflow/go/core/framework/tensor_shape_go_proto\xf8\x01\x01\x62\x06proto3')
24 )
29 _TENSORSHAPEPROTO_DIM = _descriptor.Descriptor(
30 name='Dim',
31 full_name='tensorflow.TensorShapeProto.Dim',
32 filename=None,
33 file=DESCRIPTOR,
34 containing_type=None,
35 fields=[
---> 36 _descriptor.FieldDescriptor(
37 name='size', full_name='tensorflow.TensorShapeProto.Dim.size', index=0,
38 number=1, type=3, cpp_type=2, label=1,
39 has_default_value=False, default_value=0,
40 message_type=None, enum_type=None, containing_type=None,
41 is_extension=False, extension_scope=None,
42 serialized_options=None, file=DESCRIPTOR),
43 _descriptor.FieldDescriptor(
44 name='name', full_name='tensorflow.TensorShapeProto.Dim.name', index=1,
45 number=2, type=9, cpp_type=9, label=1,
46 has_default_value=False, default_value=_b("").decode('utf-8'),
47 message_type=None, enum_type=None, containing_type=None,
48 is_extension=False, extension_scope=None,
49 serialized_options=None, file=DESCRIPTOR),
50 ],
51 extensions=[
52 ],
53 nested_types=[],
54 enum_types=[
55 ],
56 serialized_options=None,
57 is_extendable=False,
58 syntax='proto3',
59 extension_ranges=[],
60 oneofs=[
61 ],
62 serialized_start=149,
63 serialized_end=182,
64 )
66 _TENSORSHAPEPROTO = _descriptor.Descriptor(
67 name='TensorShapeProto',
68 full_name='tensorflow.TensorShapeProto',
(...)
100 serialized_end=182,
101 )
103 _TENSORSHAPEPROTO_DIM.containing_type = _TENSORSHAPEPROTO
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/google/protobuf/descriptor.py:675, in FieldDescriptor.__new__(cls, name, full_name, index, number, type, cpp_type, label, default_value, message_type, enum_type, containing_type, is_extension, extension_scope, options, serialized_options, has_default_value, containing_oneof, json_name, file, create_key)
652 def __new__(
653 cls,
654 name,
(...)
673 create_key=None,
674 ): # pylint: disable=redefined-builtin
--> 675 _message.Message._CheckCalledFromGeneratedFile()
676 if is_extension:
677 return _message.default_pool.FindExtensionByName(full_name)
TypeError: Descriptors cannot be created directly.
If this call came from a _pb2.py file, your generated code is out of date and must be regenerated with protoc >= 3.19.0.
If you cannot immediately regenerate your protos, some other possible workarounds are:
1. Downgrade the protobuf package to 3.20.x or lower.
2. Set PROTOCOL_BUFFERS_PYTHON_IMPLEMENTATION=python (but this will use pure-Python parsing and will be much slower).
More information: https://developers.google.com/protocol-buffers/docs/news/2022-05-06#python-updates
fpath_NODE = HOME_DIR + "models_NODE_3/"
# Create the analysis object:
analysis_NODE = Analysis_TT(
run_name = "NODE_3_3bff",
filepath = fpath_NODE)
analysis_NODE.plot_trial_io(num_trials = 2)
---------------------------------------------------------------------------
NameError Traceback (most recent call last)
Cell In[23], line 3
1 fpath_NODE = HOME_DIR + "models_NODE_3/"
2 # Create the analysis object:
----> 3 analysis_NODE = Analysis_TT(
4 run_name = "NODE_3_3bff",
5 filepath = fpath_NODE)
7 analysis_NODE.plot_trial_io(num_trials = 2)
NameError: name 'Analysis_TT' is not defined
There are also useful data visualization functions, such as visualizing a scree plot of the latent activity.
A scree plot shows the % of variance in the highest principle component dimensions. From this plot, we can see that the GRU has the majority of its variance in the first 3 PCs, but significant variance remains in the lower PCs!
analysis_GRU_128.plot_scree()
---------------------------------------------------------------------------
NameError Traceback (most recent call last)
Cell In[24], line 1
----> 1 analysis_GRU_128.plot_scree()
NameError: name 'analysis_GRU_128' is not defined
Importantly, the analysis object also provides functions that give access to the raw latent activity, predicted outputs, etc. of the trained models! All of these functions accept a “phase” variable that designates whether to return the training and/or validation datasets. These functions are:
get_latents(): Returns latent activity of the trained modelget_inputs(): Returns the inputs to the model (for 3BFF, the input pulses)get_model_output(): Returns a dict that contains all model outputs:controlled - the variable that the model is controlling
latents - the latent activity
actions - the output from the model (for RandomTarget only)
states - the state of the environment (for RandomTarget only)
joints - Joint angles (for RandomTarget only)
print(f"All data shape: {analysis_GRU_128.get_latents().shape}")
print(f"Train data shape: {analysis_GRU_128.get_latents(phase = 'train').shape}")
print(f"Validation data shape: {analysis_GRU_128.get_latents(phase = 'val').shape}")
---------------------------------------------------------------------------
NameError Traceback (most recent call last)
Cell In[25], line 1
----> 1 print(f"All data shape: {analysis_GRU_128.get_latents().shape}")
2 print(f"Train data shape: {analysis_GRU_128.get_latents(phase = 'train').shape}")
3 print(f"Validation data shape: {analysis_GRU_128.get_latents(phase = 'val').shape}")
NameError: name 'analysis_GRU_128' is not defined
Submit your feedback#
Show code cell source
# @title Submit your feedback
content_review(f"{feedback_prefix}_visualizing_latent_activity")
Part 2: Using affine transformations to compare latent activity#
Now that we have the latent activity for the 64D and the 128D GRU models trained on 3BFf, we can investigate how similar their latent activity is.
One problem: The models may be arbitrarily rotated, scaled, and translated relative to each other!
This means that we need to find the best “fit” between the two models that doesn’t fail when they are equivalent under an “affine” transformation (meaning a linear transformation and/or translation).
Luckily, we have a tool that can solve this problem for us! Linear regression.
In this code, we are:
Getting the latent activity from each model
Performing PCA on the latent activity (to get the dimensions ordered by their variance)
Fit a linear regression from one set of latent activity to the other.
from sklearn.linear_model import LinearRegression
from sklearn.metrics import r2_score
from sklearn.decomposition import PCA
source = analysis_GRU_128
target = analysis_NODE
# Get the latent activity from the validation phase for each model:
latents_source = source.get_latents(phase='train').detach().numpy()
latents_targ = target.get_latents(phase='train').detach().numpy()
latents_source_val = source.get_latents(phase='val').detach().numpy()
latents_targ_val = target.get_latents(phase='val').detach().numpy()
n_trials, n_timesteps, n_latent_source = latents_source.shape
n_trials, n_timesteps, n_latent_targ = latents_targ.shape
n_trials_val, n_timesteps_val, n_latent_source_val = latents_source_val.shape
n_trials_val, n_timesteps_val, n_latent_targ_val = latents_targ_val.shape
print(f"Latent shape for source model: {latents_source.shape}"
f"\nLatent shape for target model: {latents_targ.shape}")
---------------------------------------------------------------------------
NameError Traceback (most recent call last)
Cell In[27], line 5
2 from sklearn.metrics import r2_score
3 from sklearn.decomposition import PCA
----> 5 source = analysis_GRU_128
6 target = analysis_NODE
8 # Get the latent activity from the validation phase for each model:
NameError: name 'analysis_GRU_128' is not defined
# Perform PCA on both latent spaces to find axes of highest variance
pca_source = PCA()
pca_targ = PCA()
lats_source_pca = pca_source.fit_transform(latents_source.reshape(-1, n_latent_source)).reshape((n_trials, n_timesteps, -1))
lats_source_pca_val = pca_source.transform(latents_source_val.reshape(-1, n_latent_source)).reshape((n_trials, n_timesteps, -1))
lats_targ_pca = pca_targ.fit_transform(latents_targ.reshape(-1, n_latent_targ)).reshape((n_trials, n_timesteps, -1))
lats_targ_pca_val = pca_targ.transform(latents_targ_val.reshape(-1, n_latent_targ_val)).reshape((n_trials_val, n_timesteps_val, -1))
# Fit a linear regression model to predict the target latents from the source latents
reg = LinearRegression().fit(lats_source_pca.reshape(-1, n_latent_source), lats_targ_pca.reshape(-1, n_latent_targ))
# Get the R2 of the fit
preds = reg.predict(lats_source_pca_val.reshape(-1, n_latent_source_val))
r2s = r2_score(lats_targ_pca_val.reshape((-1, n_latent_targ_val)), preds, multioutput = "raw_values")
r2_var = r2_score(lats_targ_pca_val.reshape((-1, n_latent_targ_val)), preds, multioutput = "variance_weighted")
print(f"R2 of linear regression fit: {r2s}")
print(f"Variance-weighted R2 of linear regression fit: {r2_var}")
---------------------------------------------------------------------------
NameError Traceback (most recent call last)
Cell In[28], line 4
2 pca_source = PCA()
3 pca_targ = PCA()
----> 4 lats_source_pca = pca_source.fit_transform(latents_source.reshape(-1, n_latent_source)).reshape((n_trials, n_timesteps, -1))
5 lats_source_pca_val = pca_source.transform(latents_source_val.reshape(-1, n_latent_source)).reshape((n_trials, n_timesteps, -1))
7 lats_targ_pca = pca_targ.fit_transform(latents_targ.reshape(-1, n_latent_targ)).reshape((n_trials, n_timesteps, -1))
NameError: name 'latents_source' is not defined
So, the variance weighted R2 from the source to the target is ~0.93.
Importantly, we had to pick a “direction” to compute this R2 value. What happens if we switch the source and targets?
❓❓❓
Try reversing the direction (the source and targets) and see how well the model fits!
❓❓❓
One final tool that is provided to you is the comparison object, which makes many of these direct comparisons within the object itself. Here is one example visualization that shows how similar the latent activities of two example trials are for these two models!
This function has the affine transformation “built-in,” so you don’t need this to show what your R2 value above looks like in the first 3 PCs!
from ctd.comparison.comparison import Comparison
comp = Comparison()
comp.load_analysis(analysis_GRU_128, reference_analysis=True)
comp.load_analysis(analysis_NODE)
comp.plot_trials_3d_reference(num_trials=2)
---------------------------------------------------------------------------
TypeError Traceback (most recent call last)
Cell In[29], line 1
----> 1 from ctd.comparison.comparison import Comparison
2 comp = Comparison()
3 comp.load_analysis(analysis_GRU_128, reference_analysis=True)
File ~/work/NeuroAI_Course/NeuroAI_Course/projects/project-notebooks/ComputationThruDynamicsBenchmark/ctd/comparison/comparison.py:4
2 import numpy as np
3 import torch
----> 4 from DSA import DSA
5 from scipy.spatial import procrustes
6 from sklearn.cross_decomposition import CCA
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/DSA/__init__.py:3
1 __version__ = "2.0.0"
----> 3 from DSA.dsa import DSA, ControllabilitySimilarityTransformDistConfig, GeneralizedDSA, InputDSA, SimilarityTransformDistConfig
4 from DSA.dsa import DefaultDMDConfig as DMDConfig
5 from DSA.dsa import pyKoopmanDMDConfig, SubspaceDMDcConfig, DMDcConfig
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/DSA/dsa.py:5
3 from DSA.dmdc import DMDc as DefaultDMDc
4 from DSA.subspace_dmdc import SubspaceDMDc
----> 5 from DSA.simdist import SimilarityTransformDist
6 from typing import Literal
7 import torch
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/DSA/simdist.py:9
6 import torch.nn.utils.parametrize as parametrize
7 import warnings
----> 9 from ot import dist, emd, emd2, sinkhorn2
11 try:
12 from .dmd import DMD
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/ot/__init__.py:20
1 """
2 .. warning::
3 The list of automatically imported sub-modules is as follows:
(...)
11 - :any:`ot.plot` : depends on :code:`matplotlib`
12 """
14 # Author: Remi Flamary <remi.flamary@unice.fr>
15 # Nicolas Courty <ncourty@irisa.fr>
16 #
17 # License: MIT License
18
19 # All submodules and packages
---> 20 from . import lp
21 from . import bregman
22 from . import optim
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/ot/lp/__init__.py:11
2 """
3 Solvers for the original linear program OT problem.
4
5 """
7 # Author: Remi Flamary <remi.flamary@unice.fr>
8 #
9 # License: MIT License
---> 11 from .dmmot import dmmot_monge_1dgrid_loss, dmmot_monge_1dgrid_optimize
12 from ._network_simplex import emd, emd2
13 from ._barycenter_solvers import (
14 barycenter,
15 free_support_barycenter,
(...)
19 NorthWestMMGluing,
20 )
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/ot/lp/dmmot.py:12
6 # Author: Ronak Mehta <ronakrm@cs.wisc.edu>
7 # Xizheng Yu <xyu354@wisc.edu>
8 #
9 # License: MIT License
11 import numpy as np
---> 12 from ..backend import get_backend
15 def dist_monge_max_min(i):
16 r"""
17 A tensor :math:c is Monge if for all valid :math:i_1, \ldots i_d and
18 :math:j_1, \ldots, j_d,
(...)
54 Workshop on Discrete Algorithms.
55 """
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/ot/backend.py:148
146 if not os.environ.get(DISABLE_TF_KEY, False):
147 try:
--> 148 import tensorflow as tf
149 import tensorflow.experimental.numpy as tnp
151 tf_type = tf.Tensor
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/tensorflow/__init__.py:37
34 import sys as _sys
35 import typing as _typing
---> 37 from tensorflow.python.tools import module_util as _module_util
38 from tensorflow.python.util.lazy_loader import LazyLoader as _LazyLoader
40 # Make sure code inside the TensorFlow codebase can use tf2.enabled() at import.
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/tensorflow/python/__init__.py:37
29 # We aim to keep this file minimal and ideally remove completely.
30 # If you are adding a new file with @tf_export decorators,
31 # import it in modules_with_exports.py instead.
32
33 # go/tf-wildcard-import
34 # pylint: disable=wildcard-import,g-bad-import-order,g-import-not-at-top
36 from tensorflow.python import pywrap_tensorflow as _pywrap_tensorflow
---> 37 from tensorflow.python.eager import context
39 # pylint: enable=wildcard-import
40
41 # Bring in subpackages.
42 from tensorflow.python import data
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/tensorflow/python/eager/context.py:29
26 import numpy as np
27 import six
---> 29 from tensorflow.core.framework import function_pb2
30 from tensorflow.core.protobuf import config_pb2
31 from tensorflow.core.protobuf import coordination_config_pb2
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/tensorflow/core/framework/function_pb2.py:16
11 # @@protoc_insertion_point(imports)
13 _sym_db = _symbol_database.Default()
---> 16 from tensorflow.core.framework import attr_value_pb2 as tensorflow_dot_core_dot_framework_dot_attr__value__pb2
17 from tensorflow.core.framework import node_def_pb2 as tensorflow_dot_core_dot_framework_dot_node__def__pb2
18 from tensorflow.core.framework import op_def_pb2 as tensorflow_dot_core_dot_framework_dot_op__def__pb2
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/tensorflow/core/framework/attr_value_pb2.py:16
11 # @@protoc_insertion_point(imports)
13 _sym_db = _symbol_database.Default()
---> 16 from tensorflow.core.framework import tensor_pb2 as tensorflow_dot_core_dot_framework_dot_tensor__pb2
17 from tensorflow.core.framework import tensor_shape_pb2 as tensorflow_dot_core_dot_framework_dot_tensor__shape__pb2
18 from tensorflow.core.framework import types_pb2 as tensorflow_dot_core_dot_framework_dot_types__pb2
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/tensorflow/core/framework/tensor_pb2.py:16
11 # @@protoc_insertion_point(imports)
13 _sym_db = _symbol_database.Default()
---> 16 from tensorflow.core.framework import resource_handle_pb2 as tensorflow_dot_core_dot_framework_dot_resource__handle__pb2
17 from tensorflow.core.framework import tensor_shape_pb2 as tensorflow_dot_core_dot_framework_dot_tensor__shape__pb2
18 from tensorflow.core.framework import types_pb2 as tensorflow_dot_core_dot_framework_dot_types__pb2
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/tensorflow/core/framework/resource_handle_pb2.py:16
11 # @@protoc_insertion_point(imports)
13 _sym_db = _symbol_database.Default()
---> 16 from tensorflow.core.framework import tensor_shape_pb2 as tensorflow_dot_core_dot_framework_dot_tensor__shape__pb2
17 from tensorflow.core.framework import types_pb2 as tensorflow_dot_core_dot_framework_dot_types__pb2
20 DESCRIPTOR = _descriptor.FileDescriptor(
21 name='tensorflow/core/framework/resource_handle.proto',
22 package='tensorflow',
(...)
26 ,
27 dependencies=[tensorflow_dot_core_dot_framework_dot_tensor__shape__pb2.DESCRIPTOR,tensorflow_dot_core_dot_framework_dot_types__pb2.DESCRIPTOR,])
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/tensorflow/core/framework/tensor_shape_pb2.py:36
13 _sym_db = _symbol_database.Default()
18 DESCRIPTOR = _descriptor.FileDescriptor(
19 name='tensorflow/core/framework/tensor_shape.proto',
20 package='tensorflow',
(...)
23 serialized_pb=_b('\n,tensorflow/core/framework/tensor_shape.proto\x12\ntensorflow\"z\n\x10TensorShapeProto\x12-\n\x03\x64im\x18\x02 \x03(\x0b\x32 .tensorflow.TensorShapeProto.Dim\x12\x14\n\x0cunknown_rank\x18\x03 \x01(\x08\x1a!\n\x03\x44im\x12\x0c\n\x04size\x18\x01 \x01(\x03\x12\x0c\n\x04name\x18\x02 \x01(\tB\x87\x01\n\x18org.tensorflow.frameworkB\x11TensorShapeProtosP\x01ZSgithub.com/tensorflow/tensorflow/tensorflow/go/core/framework/tensor_shape_go_proto\xf8\x01\x01\x62\x06proto3')
24 )
29 _TENSORSHAPEPROTO_DIM = _descriptor.Descriptor(
30 name='Dim',
31 full_name='tensorflow.TensorShapeProto.Dim',
32 filename=None,
33 file=DESCRIPTOR,
34 containing_type=None,
35 fields=[
---> 36 _descriptor.FieldDescriptor(
37 name='size', full_name='tensorflow.TensorShapeProto.Dim.size', index=0,
38 number=1, type=3, cpp_type=2, label=1,
39 has_default_value=False, default_value=0,
40 message_type=None, enum_type=None, containing_type=None,
41 is_extension=False, extension_scope=None,
42 serialized_options=None, file=DESCRIPTOR),
43 _descriptor.FieldDescriptor(
44 name='name', full_name='tensorflow.TensorShapeProto.Dim.name', index=1,
45 number=2, type=9, cpp_type=9, label=1,
46 has_default_value=False, default_value=_b("").decode('utf-8'),
47 message_type=None, enum_type=None, containing_type=None,
48 is_extension=False, extension_scope=None,
49 serialized_options=None, file=DESCRIPTOR),
50 ],
51 extensions=[
52 ],
53 nested_types=[],
54 enum_types=[
55 ],
56 serialized_options=None,
57 is_extendable=False,
58 syntax='proto3',
59 extension_ranges=[],
60 oneofs=[
61 ],
62 serialized_start=149,
63 serialized_end=182,
64 )
66 _TENSORSHAPEPROTO = _descriptor.Descriptor(
67 name='TensorShapeProto',
68 full_name='tensorflow.TensorShapeProto',
(...)
100 serialized_end=182,
101 )
103 _TENSORSHAPEPROTO_DIM.containing_type = _TENSORSHAPEPROTO
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/google/protobuf/descriptor.py:675, in FieldDescriptor.__new__(cls, name, full_name, index, number, type, cpp_type, label, default_value, message_type, enum_type, containing_type, is_extension, extension_scope, options, serialized_options, has_default_value, containing_oneof, json_name, file, create_key)
652 def __new__(
653 cls,
654 name,
(...)
673 create_key=None,
674 ): # pylint: disable=redefined-builtin
--> 675 _message.Message._CheckCalledFromGeneratedFile()
676 if is_extension:
677 return _message.default_pool.FindExtensionByName(full_name)
TypeError: Descriptors cannot be created directly.
If this call came from a _pb2.py file, your generated code is out of date and must be regenerated with protoc >= 3.19.0.
If you cannot immediately regenerate your protos, some other possible workarounds are:
1. Downgrade the protobuf package to 3.20.x or lower.
2. Set PROTOCOL_BUFFERS_PYTHON_IMPLEMENTATION=python (but this will use pure-Python parsing and will be much slower).
More information: https://developers.google.com/protocol-buffers/docs/news/2022-05-06#python-updates
Submit your feedback#
Show code cell source
# @title Submit your feedback
content_review(f"{feedback_prefix}_using_affine_transformations_to_compare_latent_activity")
Part 3: Fixed-point finding#
Finally, we can use fixed-point finding to inspect the linearized dynamics of the trained model.
What are fixed-points?
Fixed points are points in the dynamics for which the flow field is zero, meaning that points at that location do not move.
The fixed point structure for the 3BFF task was first shown in the original Sussillo and Barack paper.

We can see that the fixed-points are at the vertices of the cube above, drawing the activity towards them and keeping it there until an input pushes it out!
We use a modified version of a fixed point finder released by Golub et al. 2018 to search the flow field for these zero points.
❓❓❓
Try changing some of these parameters:
How quickly are the fixed-points found in the model?
How many initializations are needed to find the fixed points?
Do the stability properties tell us anything about the underlying computation?
❓❓❓
Importantly from Driscol et al. 2022, we know that changes in the inputs can have large effects on the fixed point architecture, so we’re going to set the inputs to zero in this optimization.
import torch
import contextlib
import io
with contextlib.redirect_stdout(io.StringIO()): #to suppress output
fps = analysis_GRU_128.plot_fps(
inputs= torch.zeros(3),
n_inits=1024,
learning_rate=1e-3,
noise_scale=0.0,
max_iters=20000,
seed=0,
compute_jacobians=True,
q_thresh=1e-5,
)
---------------------------------------------------------------------------
NameError Traceback (most recent call last)
Cell In[31], line 6
3 import io
5 with contextlib.redirect_stdout(io.StringIO()): #to suppress output
----> 6 fps = analysis_GRU_128.plot_fps(
7 inputs= torch.zeros(3),
8 n_inits=1024,
9 learning_rate=1e-3,
10 noise_scale=0.0,
11 max_iters=20000,
12 seed=0,
13 compute_jacobians=True,
14 q_thresh=1e-5,
15 )
NameError: name 'analysis_GRU_128' is not defined
import matplotlib.pyplot as plt
q_thesh = 1e-6
q_vals = fps.qstar
x_star = fps.xstar[q_vals < q_thesh]
fig = plt.figure(figsize=(10, 10))
ax = fig.add_subplot(111, projection='3d')
ax.scatter(x_star[:, 0], x_star[:, 1], x_star[:, 2], c='r', marker='o')
fig.show()
---------------------------------------------------------------------------
NameError Traceback (most recent call last)
Cell In[32], line 3
1 import matplotlib.pyplot as plt
2 q_thesh = 1e-6
----> 3 q_vals = fps.qstar
4 x_star = fps.xstar[q_vals < q_thesh]
5 fig = plt.figure(figsize=(10, 10))
NameError: name 'fps' is not defined
❓❓❓
What can you find out about the FPs of the trained models? Can you modify the FP finding to get more interpretable results?
What can we learn about the computational solution built in this 3BFF network from these fixed-point architectures?
❓❓❓
Submit your feedback#
Show code cell source
# @title Submit your feedback
content_review(f"{feedback_prefix}_fixed_point_finding")
Section 6: Introducing the Random Target task#
Now that we’ve developed intuition on a simple, well-understood task, let’s move up the ladder of complexity!
The second task is a random-target reaching task performed by an RNN controlling a 2-joint musculoskeletal model of an arm actuated by 6 Mujoco muscles. This environment was built using MotorNet, a package developed by Oli Codol et al. that provides environments for training RNNs to control biomechanical models!
Here is a short clip of what this task looks like when performed by a well-trained model:

Behaviorally, the task has the following structure:
A random initial hand position is sampled from a range of reachable locations; the model is instructed to maintain that hand position.
A random target position is chosen from the range of reachable locations and fed to the model.
After a random delay period, a go-cue is fed to the model, which prompts the model to generate muscle activations that drive the hand to the target location.
On 20% of trials, the go-cue is never supplied (“catch” trials)
On 50% of trials, a randomly directed bump perturbation (5-10 N, 150-300 ms duration) is applied to the hand.
50% of these bumps occur in a small window after the go-cue
50% of these bumps occur at a random time in the trial
The model is trained to:
Minimize the MSE between the hand position and the desired hand position
Minimize the squared muscle activation
with each loss term being weighted by a scalar.
from ctd.task_modeling.task_env.task_env import RandomTarget
from motornet.effector import RigidTendonArm26
from motornet.muscle import MujocoHillMuscle
# Create the analysis object:
rt_task_env = RandomTarget(effector = RigidTendonArm26(muscle = MujocoHillMuscle()))
⏳⏳⏳
Now, to train the model! We use the same procedure as the 3BFF above; however, this model will take a bit longer to train, as of the serial nature of this task, the parallelization allowed by GPUs doesn’t help speed up our training!
⏳⏳⏳
from ctd.task_modeling.model.rnn import GRU_RNN
from ctd.task_modeling.datamodule.task_datamodule import TaskDataModule
from ctd.task_modeling.task_wrapper.task_wrapper import TaskTrainedWrapper
from pytorch_lightning import Trainer
# Step 1: Instantiate the model
rnn = GRU_RNN(latent_size = 128) # Look in ctd/task_modeling/models for alternative choices!
# Step 2: Instantiate the task environment
task_env = rt_task_env
# Step 3: Instantiate the task datamodule
task_datamodule = TaskDataModule(task_env, n_samples = 1000, batch_size = 256)
# Step 4: Instantiate the task wrapper
task_wrapper = TaskTrainedWrapper(learning_rate=1e-3, weight_decay = 1e-8)
# Step 5: Initialize the model with the input and output sizes
rnn.init_model(
input_size = task_env.observation_space.shape[0] + task_env.context_inputs.shape[0],
output_size = task_env.action_space.shape[0]
)
# Step 6: Set the environment and model in the task wrapper
task_wrapper.set_environment(task_env)
task_wrapper.set_model(rnn)
# Step 7: Define the PyTorch Lightning Trainer object (put `enable_progress_bar=True` to observe training progress)
trainer = Trainer(accelerator= "cpu",max_epochs=500,enable_progress_bar=False)
# Step 8: Fit the model
trainer.fit(task_wrapper, task_datamodule)
---------------------------------------------------------------------------
TypeError Traceback (most recent call last)
Cell In[35], line 32
29 trainer = Trainer(accelerator= "cpu",max_epochs=500,enable_progress_bar=False)
31 # Step 8: Fit the model
---> 32 trainer.fit(task_wrapper, task_datamodule)
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/pytorch_lightning/trainer/trainer.py:544, in Trainer.fit(self, model, train_dataloaders, val_dataloaders, datamodule, ckpt_path)
542 self.state.status = TrainerStatus.RUNNING
543 self.training = True
--> 544 call._call_and_handle_interrupt(
545 self, self._fit_impl, model, train_dataloaders, val_dataloaders, datamodule, ckpt_path
546 )
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/pytorch_lightning/trainer/call.py:44, in _call_and_handle_interrupt(trainer, trainer_fn, *args, **kwargs)
42 if trainer.strategy.launcher is not None:
43 return trainer.strategy.launcher.launch(trainer_fn, *args, trainer=trainer, **kwargs)
---> 44 return trainer_fn(*args, **kwargs)
46 except _TunerExitException:
47 _call_teardown_hook(trainer)
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/pytorch_lightning/trainer/trainer.py:580, in Trainer._fit_impl(self, model, train_dataloaders, val_dataloaders, datamodule, ckpt_path)
573 assert self.state.fn is not None
574 ckpt_path = self._checkpoint_connector._select_ckpt_path(
575 self.state.fn,
576 ckpt_path,
577 model_provided=True,
578 model_connected=self.lightning_module is not None,
579 )
--> 580 self._run(model, ckpt_path=ckpt_path)
582 assert self.state.stopped
583 self.training = False
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/pytorch_lightning/trainer/trainer.py:950, in Trainer._run(self, model, ckpt_path)
947 self.strategy.setup_environment()
948 self.__setup_profiler()
--> 950 call._call_setup_hook(self) # allow user to setup lightning_module in accelerator environment
952 # check if we should delay restoring checkpoint till later
953 if not self.strategy.restore_checkpoint_after_setup:
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/pytorch_lightning/trainer/call.py:86, in _call_setup_hook(trainer)
84 # Trigger lazy creation of experiment in loggers so loggers have their metadata available
85 for logger in trainer.loggers:
---> 86 if hasattr(logger, "experiment"):
87 _ = logger.experiment
89 trainer.strategy.barrier("pre_setup")
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/lightning_fabric/loggers/logger.py:118, in rank_zero_experiment.<locals>.experiment(self)
116 if rank_zero_only.rank > 0:
117 return _DummyExperiment()
--> 118 return fn(self)
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/lightning_fabric/loggers/tensorboard.py:187, in TensorBoardLogger.experiment(self)
184 self._fs.makedirs(self.root_dir, exist_ok=True)
186 if _TENSORBOARD_AVAILABLE:
--> 187 from torch.utils.tensorboard import SummaryWriter
188 else:
189 from tensorboardX import SummaryWriter # type: ignore[no-redef]
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/torch/utils/tensorboard/__init__.py:12
9 del Version
10 del tensorboard
---> 12 from .writer import FileWriter, SummaryWriter # noqa: F401
13 from tensorboard.summary.writer.record_writer import RecordWriter # noqa: F401
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/torch/utils/tensorboard/writer.py:10
7 import torch
9 from tensorboard.compat import tf
---> 10 from tensorboard.compat.proto import event_pb2
11 from tensorboard.compat.proto.event_pb2 import Event, SessionLog
12 from tensorboard.plugins.projector.projector_config_pb2 import ProjectorConfig
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/tensorboard/compat/proto/event_pb2.py:17
12 # @@protoc_insertion_point(imports)
14 _sym_db = _symbol_database.Default()
---> 17 from tensorboard.compat.proto import summary_pb2 as tensorboard_dot_compat_dot_proto_dot_summary__pb2
20 DESCRIPTOR = _descriptor.FileDescriptor(
21 name='tensorboard/compat/proto/event.proto',
22 package='tensorboard',
(...)
26 ,
27 dependencies=[tensorboard_dot_compat_dot_proto_dot_summary__pb2.DESCRIPTOR,])
29 _WORKERHEALTH = _descriptor.EnumDescriptor(
30 name='WorkerHealth',
31 full_name='tensorboard.WorkerHealth',
(...)
55 serialized_end=1319,
56 )
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/tensorboard/compat/proto/summary_pb2.py:17
12 # @@protoc_insertion_point(imports)
14 _sym_db = _symbol_database.Default()
---> 17 from tensorboard.compat.proto import tensor_pb2 as tensorboard_dot_compat_dot_proto_dot_tensor__pb2
20 DESCRIPTOR = _descriptor.FileDescriptor(
21 name='tensorboard/compat/proto/summary.proto',
22 package='tensorboard',
(...)
26 ,
27 dependencies=[tensorboard_dot_compat_dot_proto_dot_tensor__pb2.DESCRIPTOR,])
29 _DATACLASS = _descriptor.EnumDescriptor(
30 name='DataClass',
31 full_name='tensorboard.DataClass',
(...)
55 serialized_end=1228,
56 )
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/tensorboard/compat/proto/tensor_pb2.py:16
11 # @@protoc_insertion_point(imports)
13 _sym_db = _symbol_database.Default()
---> 16 from tensorboard.compat.proto import resource_handle_pb2 as tensorboard_dot_compat_dot_proto_dot_resource__handle__pb2
17 from tensorboard.compat.proto import tensor_shape_pb2 as tensorboard_dot_compat_dot_proto_dot_tensor__shape__pb2
18 from tensorboard.compat.proto import types_pb2 as tensorboard_dot_compat_dot_proto_dot_types__pb2
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/tensorboard/compat/proto/resource_handle_pb2.py:16
11 # @@protoc_insertion_point(imports)
13 _sym_db = _symbol_database.Default()
---> 16 from tensorboard.compat.proto import tensor_shape_pb2 as tensorboard_dot_compat_dot_proto_dot_tensor__shape__pb2
17 from tensorboard.compat.proto import types_pb2 as tensorboard_dot_compat_dot_proto_dot_types__pb2
20 DESCRIPTOR = _descriptor.FileDescriptor(
21 name='tensorboard/compat/proto/resource_handle.proto',
22 package='tensorboard',
(...)
26 ,
27 dependencies=[tensorboard_dot_compat_dot_proto_dot_tensor__shape__pb2.DESCRIPTOR,tensorboard_dot_compat_dot_proto_dot_types__pb2.DESCRIPTOR,])
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/tensorboard/compat/proto/tensor_shape_pb2.py:36
13 _sym_db = _symbol_database.Default()
18 DESCRIPTOR = _descriptor.FileDescriptor(
19 name='tensorboard/compat/proto/tensor_shape.proto',
20 package='tensorboard',
(...)
23 serialized_pb=_b('\n+tensorboard/compat/proto/tensor_shape.proto\x12\x0btensorboard\"{\n\x10TensorShapeProto\x12.\n\x03\x64im\x18\x02 \x03(\x0b\x32!.tensorboard.TensorShapeProto.Dim\x12\x14\n\x0cunknown_rank\x18\x03 \x01(\x08\x1a!\n\x03\x44im\x12\x0c\n\x04size\x18\x01 \x01(\x03\x12\x0c\n\x04name\x18\x02 \x01(\tB\x87\x01\n\x18org.tensorflow.frameworkB\x11TensorShapeProtosP\x01ZSgithub.com/tensorflow/tensorflow/tensorflow/go/core/framework/tensor_shape_go_proto\xf8\x01\x01\x62\x06proto3')
24 )
29 _TENSORSHAPEPROTO_DIM = _descriptor.Descriptor(
30 name='Dim',
31 full_name='tensorboard.TensorShapeProto.Dim',
32 filename=None,
33 file=DESCRIPTOR,
34 containing_type=None,
35 fields=[
---> 36 _descriptor.FieldDescriptor(
37 name='size', full_name='tensorboard.TensorShapeProto.Dim.size', index=0,
38 number=1, type=3, cpp_type=2, label=1,
39 has_default_value=False, default_value=0,
40 message_type=None, enum_type=None, containing_type=None,
41 is_extension=False, extension_scope=None,
42 serialized_options=None, file=DESCRIPTOR),
43 _descriptor.FieldDescriptor(
44 name='name', full_name='tensorboard.TensorShapeProto.Dim.name', index=1,
45 number=2, type=9, cpp_type=9, label=1,
46 has_default_value=False, default_value=_b("").decode('utf-8'),
47 message_type=None, enum_type=None, containing_type=None,
48 is_extension=False, extension_scope=None,
49 serialized_options=None, file=DESCRIPTOR),
50 ],
51 extensions=[
52 ],
53 nested_types=[],
54 enum_types=[
55 ],
56 serialized_options=None,
57 is_extendable=False,
58 syntax='proto3',
59 extension_ranges=[],
60 oneofs=[
61 ],
62 serialized_start=150,
63 serialized_end=183,
64 )
66 _TENSORSHAPEPROTO = _descriptor.Descriptor(
67 name='TensorShapeProto',
68 full_name='tensorboard.TensorShapeProto',
(...)
100 serialized_end=183,
101 )
103 _TENSORSHAPEPROTO_DIM.containing_type = _TENSORSHAPEPROTO
File /opt/hostedtoolcache/Python/3.10.19/x64/lib/python3.10/site-packages/google/protobuf/descriptor.py:675, in FieldDescriptor.__new__(cls, name, full_name, index, number, type, cpp_type, label, default_value, message_type, enum_type, containing_type, is_extension, extension_scope, options, serialized_options, has_default_value, containing_oneof, json_name, file, create_key)
652 def __new__(
653 cls,
654 name,
(...)
673 create_key=None,
674 ): # pylint: disable=redefined-builtin
--> 675 _message.Message._CheckCalledFromGeneratedFile()
676 if is_extension:
677 return _message.default_pool.FindExtensionByName(full_name)
TypeError: Descriptors cannot be created directly.
If this call came from a _pb2.py file, your generated code is out of date and must be regenerated with protoc >= 3.19.0.
If you cannot immediately regenerate your protos, some other possible workarounds are:
1. Downgrade the protobuf package to 3.20.x or lower.
2. Set PROTOCOL_BUFFERS_PYTHON_IMPLEMENTATION=python (but this will use pure-Python parsing and will be much slower).
More information: https://developers.google.com/protocol-buffers/docs/news/2022-05-06#python-updates
Importantly, this task is distinct from the previous two tasks because the outputs of the model affect the subsequent inputs!

Visualizing the latent dynamics of models trained on MotorNet tasks, we can see that there are complex features in the state space, but we’ll leave that to you to figure out what they mean!

In the later questions, we will ask you to modify the environments in MotorNet to test how well your models can generalize to new tasks!
Submit your feedback#
Show code cell source
# @title Submit your feedback
content_review(f"{feedback_prefix}_introducing_the_random_target_task")
Conclusion#
That’s it!
To recap, in this tutorial, we learned:
The basics of two tasks, the Three-Bit Flip-Flop and the Random Target task.
How to train recurrent neural network models on these tasks
Methods of visualizing and quantifying differences between these task-trained models.
As you begin to extend beyond this tutorial, you will likely need to make your own environments, or modify existing environments to test the ability of models to generalize. We’ve tried to document the code-base to make this as easy as possible, but feel free to reach out if you have any questions!
Submit your feedback#
Show code cell source
# @title Submit your feedback
content_review(f"{feedback_prefix}_conclusion")

